Careers can be measured not just in years, but in impact – in the people, the science, and in commitment to a shared purpose. As Christine Williamson and Mark and Jackie Harrison prepare to retire, we reflect back on decades of change and dedication.
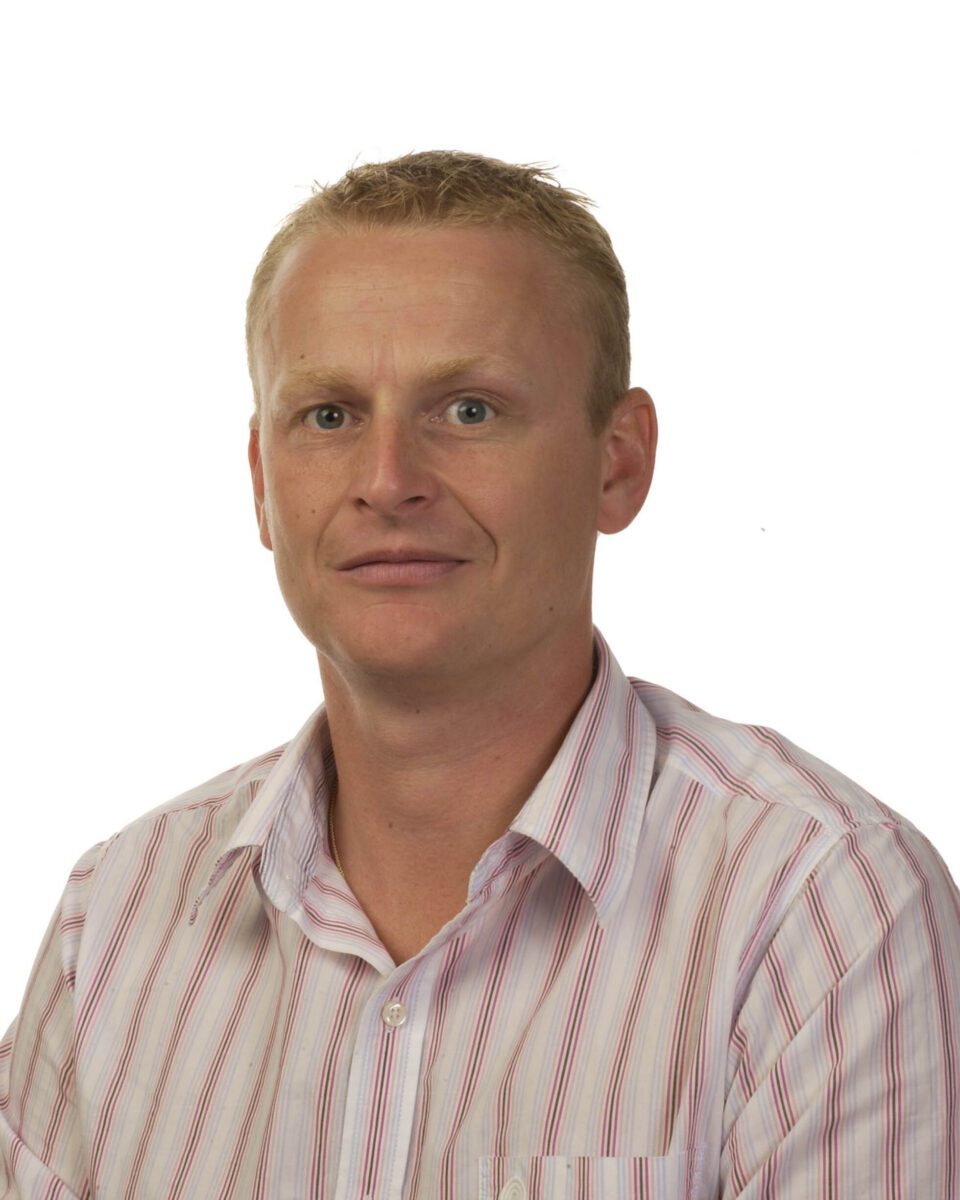
Mark Harrison: From open cages to Operations Manager
When MRC Harwell welcomed Mark Harrison in September 1989, he arrived straight from school as an assistant scientific officer—though, as he puts it, “it was really a junior animal technician.” Mark’s journey naturally evolved into his current role as Operations Manager in the Mary Lyon Centre (MLC). With years of hands-on experience behind him, the shift from working directly with animals to overseeing operations marked a significant—but logical—change. It meant stepping back from the day-to-day interactions with the mice and instead viewing the unit as a whole.
“When I started, we had no barrier in place and windows in the mouse rooms,” Mark recalls. “No gloves, face masks, or mobcaps… you could walk in wearing your own clothes under a lab coat.” There were open-top cages, the constant sound of mice, and, as he candidly puts it, “the smell was strong.” All very different from the MLC of today.
Despite all the change, one thing remained constant: his commitment.
When asked what he’s most proud of, Mark doesn’t point to a single achievement. Instead, he reflects simply on his dedication to the unit over the years—a quiet but powerful testament to a career built on consistency and reliability. That, and the claim to fame that he has met or worked with everyone who has a room named after them here at the MLC!
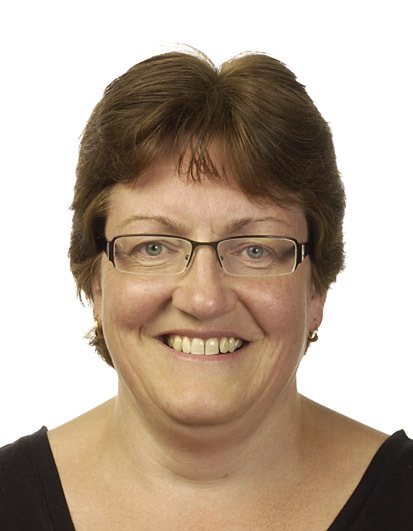



Christine Williamson: “Poacher turned gamekeeper”
Christine Williamson’s journey at MRC Harwell began in September 1992, when she joined the MRC Radiobiology Unit on a three-year postdoctoral contract. Working in Jo Peters’ team, her focus was genomic imprinting—the phenomenon that causes genes to be expressed or not, depending on whether they are inherited from the mother or father.
Even at the interview stage, Christine senses the significance of the opportunity. Meeting Bruce Cattanach, she made a point of shaking his hand, thinking that it might be her only chance to meet such a pioneering geneticist. Little did she realise, she would later be working on the imprinted region on distal mouse Chromosome 2 discovered by Bruce – work that was described as instrumental in unravelling its complex regulatory mechanisms.
Along the way, there were moments that captured the unique environment she was part of, including sitting next to Mary Lyon at a Christmas dinner and discussing what it felt like to have a building named in her honour.
Christine speaks with deep appreciation for the people she worked with, particularly Jo Peters, whose leadership and collaboration left a lasting impression. When Jo retired, she wrote to Christine thanking her for “really making those key experiments possible” and for her vital role in supporting the group.
After 18 years in research, including two first-author publications in Nature Genetics, Christine made a significant career shift. Rather than leaving MRC Harwell when Jo retired, she chose to stay—moving into the Health & Safety team in 2011. Supported by the organisation to retrain, she achieved distinctions in her qualifications and went on to become Head of Health & Safety in 2014. She reflects on the transition as perhaps being a “poacher turned gamekeeper”, but also on the benefit of the technical experience she brought to her new position.
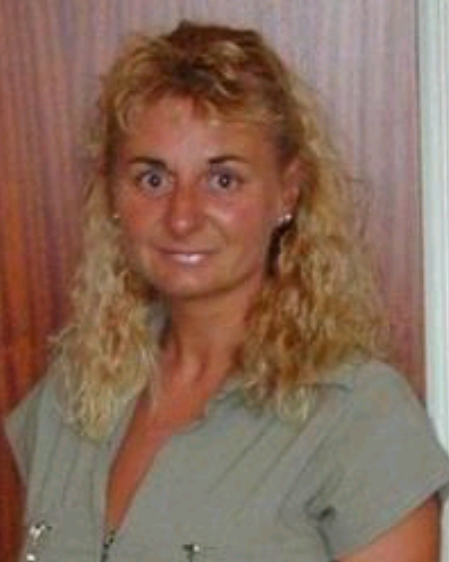
Jackie Harrison: Precision, teamwork, and pride in the details
When MRC Harwell welcomed Jackie Harrison in August 1998, she began her career as an animal technician. Today, Jackie serves as an in vivo dosing manager, following nearly 20 years as a ward manager. Her career path reflects both progression and adaptation, evolving alongside the changing structure of the organisation.
Reflecting on what kept her at MRC Harwell for so many years, Jackie points to her genuine enjoyment of the work and, just as importantly, the camaraderie within her team. That sense of shared purpose—professional yet supportive—helped create an environment where high-quality work could thrive. Together, they contributed valuable data to the scientific community, something Jackie clearly takes pride in.
And when it comes to achievements, there are many.
“It’s hard to choose,” she says, reflecting on the breadth of projects she’s been part of. Among the highlights are her contributions to sexual development studies and cancer research—areas that demanded not only technical skill but also precise timing and close collaboration with scientific teams. These projects stand out not just for their complexity, but for their importance, and Jackie is proud to have played a role in work that makes a real difference.
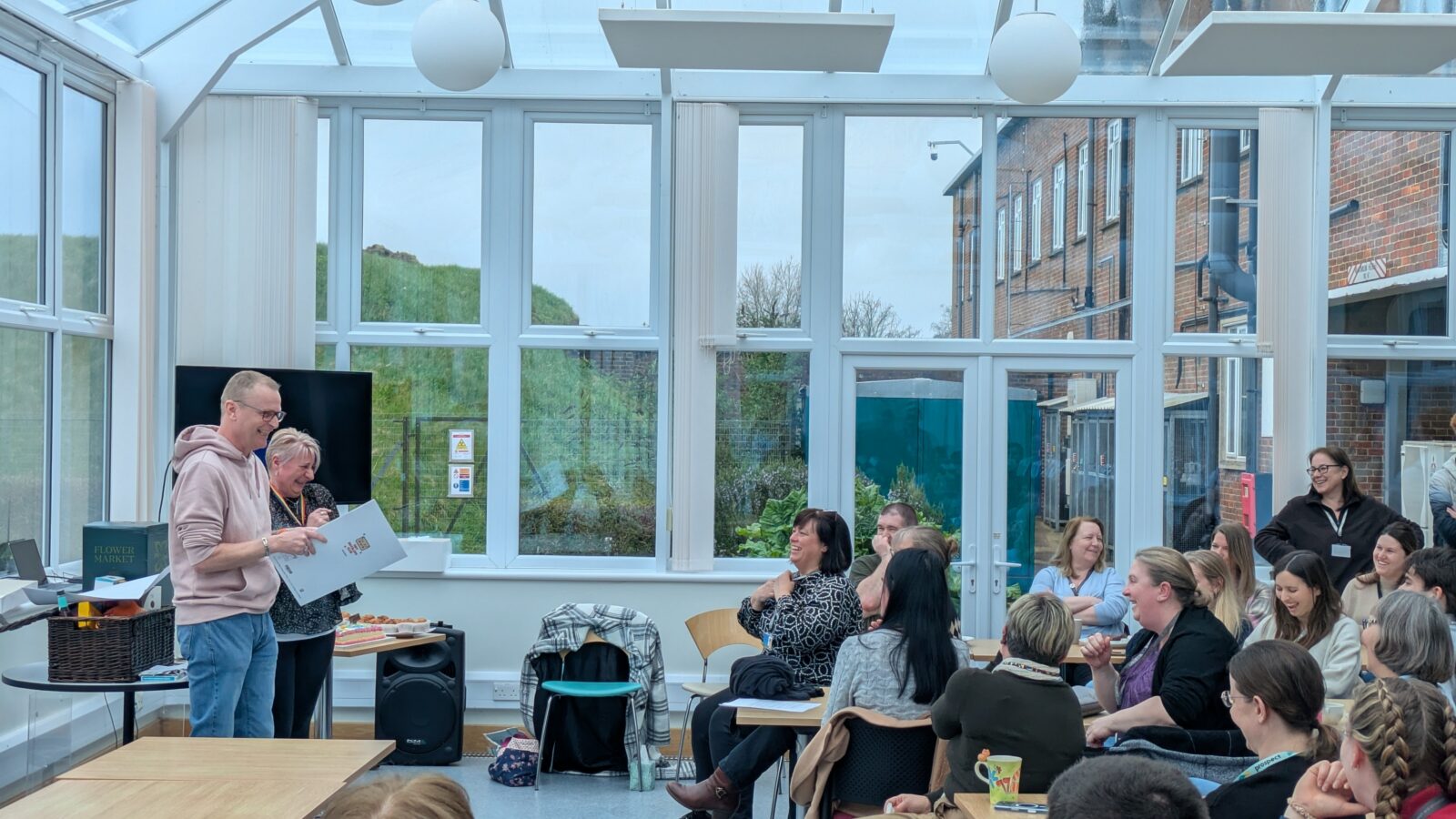



Next steps
Retirement brings a new rhythm of life. Mark and Jackie are planning to move back to Jackie’s hometown of Hereford, where Jackie wants to focus on hobbies of sewing and card making, but also do some form of voluntary work that keeps her connected with people, while Mark is looking forward to fishing on the River Wye and learning to play the bass guitar. Christine is preparing to take on the challenge of walking the 630-mile South West Coast Path in one go!
Sara Wells, Director of the MLC, spoke at Jackie and Mark’s retirement do: “Mark and Jackie are an amazing couple who have been part of my life for over twenty years, but more importantly a huge part of Harwell for over three decades. Mark’s loyalty to Harwell, and especially the Mary Lyon Centre, runs deep – he played a foundational role in the initial derivation process and was instrumental in establishing the unit itself. In Jackie, we are losing an exceptional talent, eye for animal behaviour, and a work ethic that I wish I could bottle and make everyone drink. I want them to know how much they both mean to all of us and how we will treasure the times we have shared and the many many laughs we had along the way!”
Mark Gardiner, Chief Operating Officer for the MLC, spoke at Christine’s retirement celebration: “Chris, thank you for everything: your scientific insight and your dedication to safety. I am sure that the safety culture you helped build will continue to protect colleagues long into the future. You will be missed not only as the head of safety, but as a close colleague and friend. I wish you the best of luck on your next adventure!”
Mark, Jackie, and Christine each leave behind not just years of service, but a legacy of dedication and a lasting impact on all of us. We thank them for everything they’ve contributed and wish them all the very best for the adventures ahead!