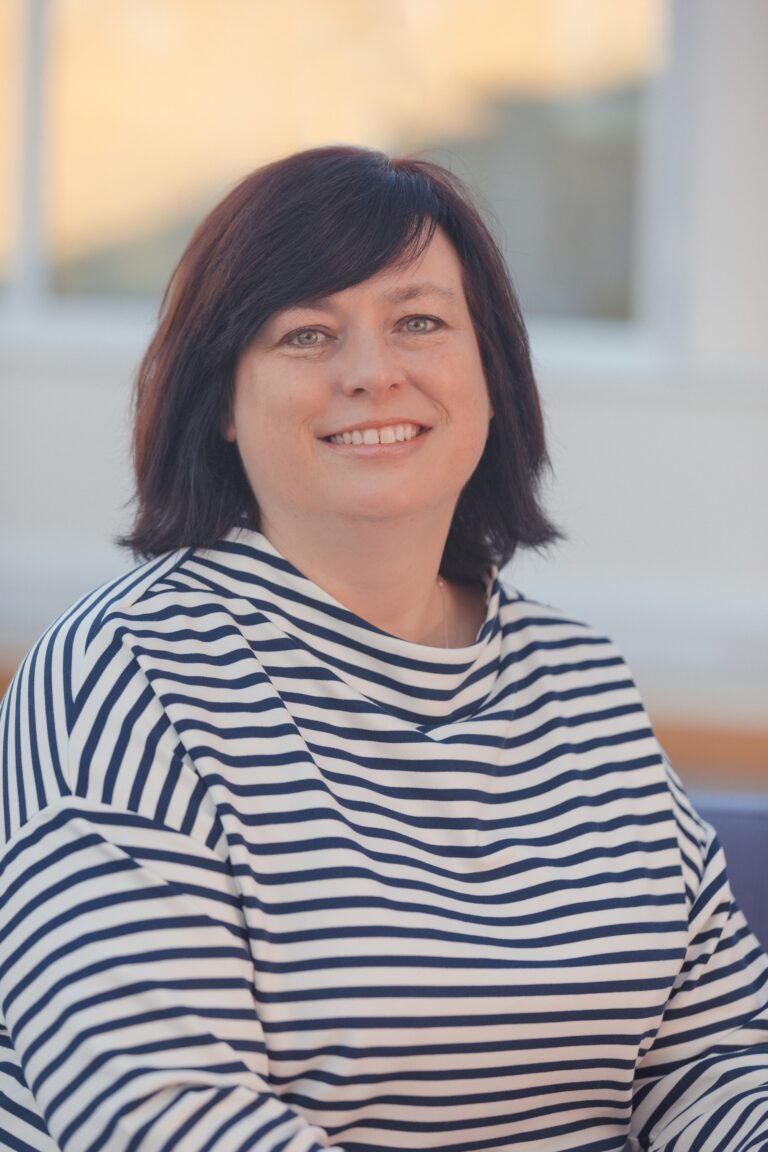
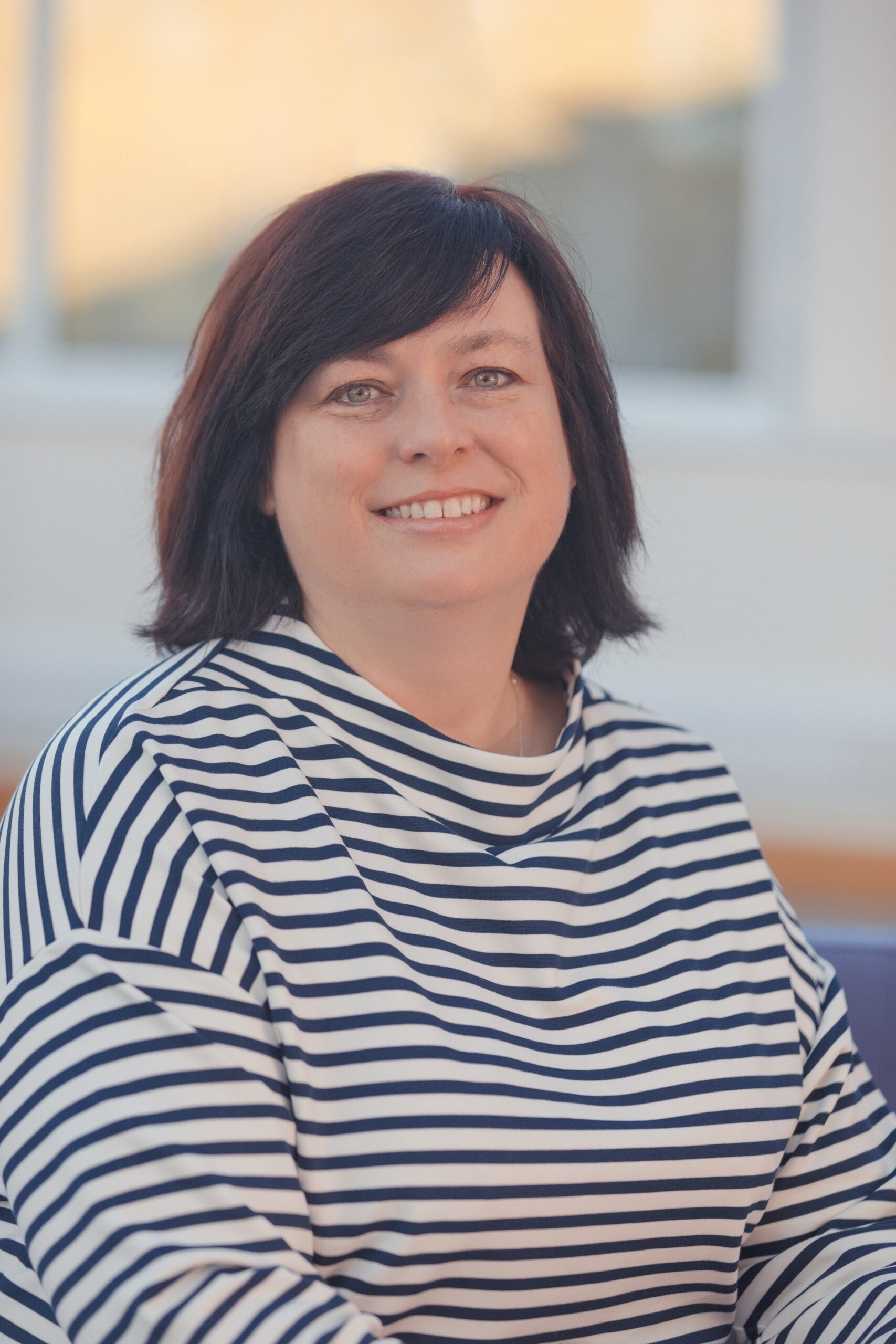
Dr Sara Wells, Director of the Mary Lyon Centre at MRC Harwell (MLC), is now the Chief Biological Resources Facility (BRF) Officer at The Francis Crick Institute. The joint appointment between the Medical Research Council and the Crick will see Sara supporting the Institute’s leadership team on strategic and practical decisions regarding the BRF while still holding the reins of the MLC.
This prestigious appointment recognises more than two decades of experience in the field as well as Sara’s outstanding track record in supporting and delivering complex research projects involving in vivo studies, and in driving the adoption of best practices in animal research throughout her career.
Sara always knew she wanted to be a scientist and her fascination with genetics began at school. The progression to a BSc in Genetics at the University of Sheffield was an obvious step forward. She went on to do a PhD on the “Expression of Neuroendocrine Genes in Transgenic Rats’ at the MRC National Institute for Medical Research but soon realised that her interest leaned more towards supporting research projects and enabling great science through the provision of the highest standards of services for the researchers.


Her experience with transgenic animals led to positions in which in vivo models were generated and used to understand fundamental biological processes with a strong focus on genetics. The perturbations of those processes that lead to disease could then be studied. Sara’s attention was always on ensuring that the scientific ideas were supported by sound experimental design and consistently controlled processes that led to the collection of good quality, reproducible data.
Sara is well known for her contagious enthusiasm for her work and her ability to find practical solutions for the most complex issues of in vivo experimental work. She often comments on how much she loves and enjoys her job.
The appointment at the Crick is a positive recognition of Sara’s competence and expertise. She said: “Supporting the BRF at the largest biomedical research institute in Europe will be a privilege and a challenge, and I am looking forward to the synergies that will no doubt develop between the two units. I expect our work to develop in ways that will be mutually beneficial, not only for the science but also in our constant commitment to the 3Rs, to ensure that we are always using the best model for a particular project, whether in vivo or ex vivo and making the most of our combined talents”. Well done boss!